statsmodels.regression.mixed_linear_model.MixedLM.EM¶
-
MixedLM.EM(fe_params, cov_re, scale, niter_em=10, hist=None)[source]¶ Run the EM algorithm from a given starting point. This is for ML (not REML), but it seems to be good enough to use for REML starting values.
Returns: fe_params : 1d ndarray
The final value of the fixed effects coefficients
cov_re : 2d ndarray
The final value of the random effects covariance matrix
scale : float
The final value of the error variance
Notes
This uses the parameterization of the likelihood
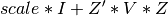 , note that this differs from the profile
likelihood used in the gradient calculations.
, note that this differs from the profile
likelihood used in the gradient calculations.
