Prediction (out of sample)¶
[1]:
%matplotlib inline
[2]:
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.api as sm
plt.rc("figure", figsize=(16, 8))
plt.rc("font", size=14)
Artificial data¶
[3]:
nsample = 50
sig = 0.25
x1 = np.linspace(0, 20, nsample)
X = np.column_stack((x1, np.sin(x1), (x1 - 5) ** 2))
X = sm.add_constant(X)
beta = [5.0, 0.5, 0.5, -0.02]
y_true = np.dot(X, beta)
y = y_true + sig * np.random.normal(size=nsample)
Estimation¶
[4]:
olsmod = sm.OLS(y, X)
olsres = olsmod.fit()
print(olsres.summary())
OLS Regression Results
==============================================================================
Dep. Variable: y R-squared: 0.986
Model: OLS Adj. R-squared: 0.985
Method: Least Squares F-statistic: 1047.
Date: Thu, 16 Apr 2026 Prob (F-statistic): 2.51e-42
Time: 11:29:46 Log-Likelihood: 5.2125
No. Observations: 50 AIC: -2.425
Df Residuals: 46 BIC: 5.223
Df Model: 3
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
const 5.0705 0.077 65.449 0.000 4.915 5.226
x1 0.4968 0.012 41.581 0.000 0.473 0.521
x2 0.4769 0.047 10.153 0.000 0.382 0.571
x3 -0.0205 0.001 -19.584 0.000 -0.023 -0.018
==============================================================================
Omnibus: 1.649 Durbin-Watson: 1.963
Prob(Omnibus): 0.438 Jarque-Bera (JB): 1.517
Skew: -0.307 Prob(JB): 0.468
Kurtosis: 2.408 Cond. No. 221.
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
In-sample prediction¶
[5]:
ypred = olsres.predict(X)
print(ypred)
[ 4.55690829 5.02940457 5.46395749 5.83457848 6.12465809 6.32969491
6.45803513 6.52950127 6.57213542 6.61759198 6.69593695 6.83070817
7.03504809 7.30954456 7.64213418 8.01008435 8.38372812 8.7313384
9.02434065 9.24200757 9.37486455 9.42624634 9.4117491 9.35666808
9.29184161 9.24858447 9.253543 9.32431711 9.46656874 9.67309155
9.92499496 10.19480777 10.45099165 10.66312275 10.80689027 10.86809069
10.8449613 10.74846988 10.60051267 10.4303169 10.26963885 10.14754637
10.08564312 10.09451913 10.17201092 10.30355694 10.46458998 10.62457352
10.75201875 10.81965814]
Create a new sample of explanatory variables Xnew, predict and plot¶
[6]:
x1n = np.linspace(20.5, 25, 10)
Xnew = np.column_stack((x1n, np.sin(x1n), (x1n - 5) ** 2))
Xnew = sm.add_constant(Xnew)
ynewpred = olsres.predict(Xnew) # predict out of sample
print(ynewpred)
[10.79475832 10.64320358 10.38369465 10.05884817 9.7247626 9.4372834
9.2383301 9.14563293 9.14839184 9.20992076]
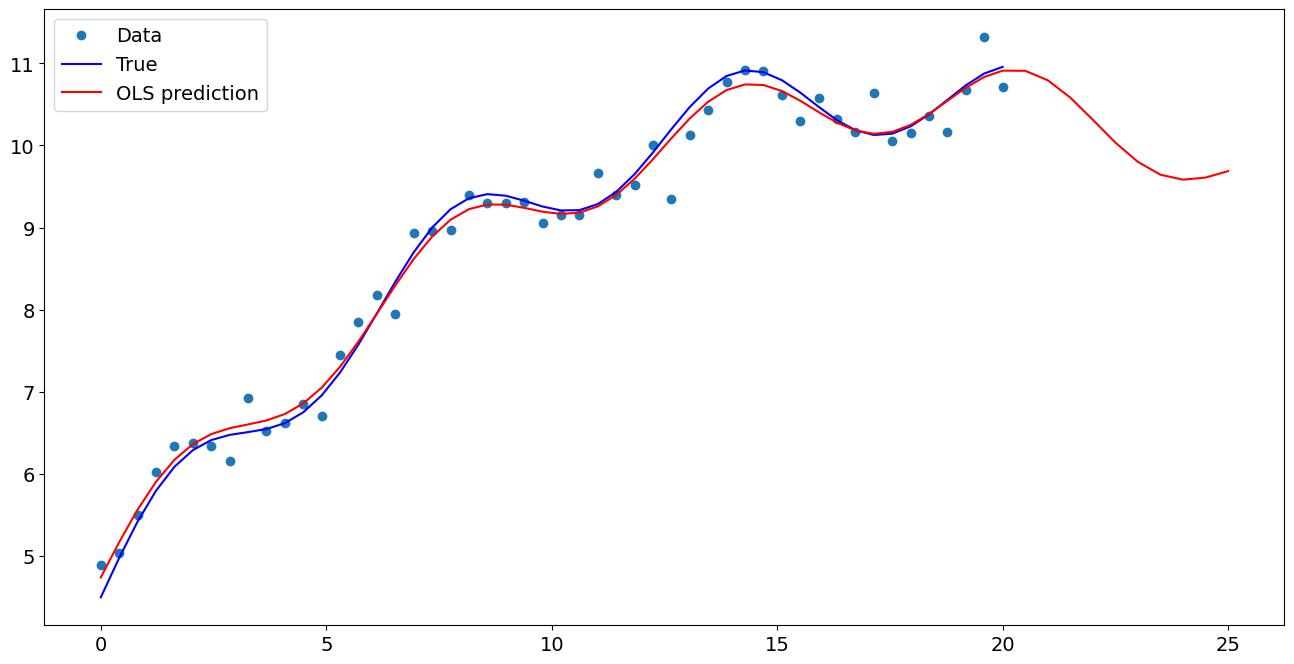
Plot comparison¶
[7]:
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.plot(x1, y, "o", label="Data")
ax.plot(x1, y_true, "b-", label="True")
ax.plot(np.hstack((x1, x1n)), np.hstack((ypred, ynewpred)), "r", label="OLS prediction")
ax.legend(loc="best")
[7]:
<matplotlib.legend.Legend at 0x7fadd410fa30>
Predicting with Formulas¶
Using formulas can make both estimation and prediction a lot easier
[8]:
from statsmodels.formula.api import ols
data = {"x1": x1, "y": y}
res = ols("y ~ x1 + np.sin(x1) + I((x1-5)**2)", data=data).fit()
We use the I to indicate use of the Identity transform. Ie., we do not want any expansion magic from using **2
[9]:
res.params
[9]:
Intercept 5.070533
x1 0.496820
np.sin(x1) 0.476862
I((x1 - 5) ** 2) -0.020545
dtype: float64
Now we only have to pass the single variable and we get the transformed right-hand side variables automatically
[10]:
res.predict(exog=dict(x1=x1n))
[10]:
0 10.794758
1 10.643204
2 10.383695
3 10.058848
4 9.724763
5 9.437283
6 9.238330
7 9.145633
8 9.148392
9 9.209921
dtype: float64